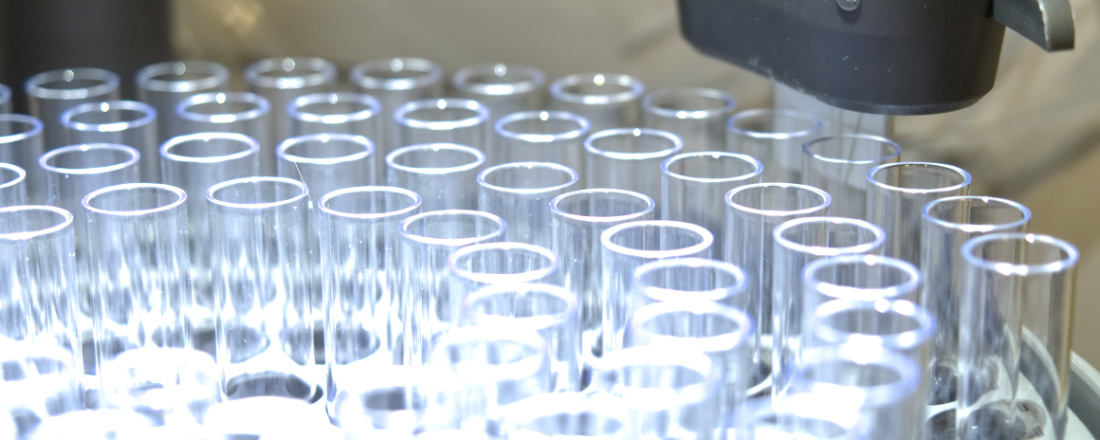
Research

Artificial Motor Proteins: Design and creation of autonomous protein motors
- together with the groups of Heiner Linke and Paul Curmi -
In an interdisciplinary and collaborative approach, we are working on the design of functional synthetic protein motors capable of moving and transducing energy, based on existing non-motor protein modules of known molecular function. Coupling the synergy of expertise in computational protein design, structural and molecular biology, and single-molecule detection from us and our collaborators from Lunds University and the University of New South Wales.
This project is funded by an ERC Synergy grant.
Selected References:
Linke H., Höcker B., Furuta K., Forde N.C., Curmi P.M.G. (2020) Synthetic biology approaches to dissecting linear motor protein function: towards the design and synthesis of artificial autonomous protein walkers. Biophys Rev [PubMed]
Computational and experimental design of protein function
The aim of this project is to use and improve computational design methodologies for the reliable construction of novel function such as catalytic activities in various protein scaffolds. Rational design is a stringent test of our understanding of biological mechanisms. And in the long run we want to be able to construct proteins with additional features for medical and industrial applications.
At the moment we are applying existing programs to well-studied reaction mechanisms in order to test their applicability. And we are working on the development of additional computational tools. An important part of this work includes the experimental test of theoretical design solutions in the lab. We are also pursuing structural studies on designed ligand-binding proteins to validate predictions in molecular detail.
Selected References:
Ferruz N., Schmidt S., Höcker B. (2022) ProtGPT2 is a deep unsupervised language model for protein design Nat Commun [PubMed]
Noske J., Kynast J.P., Lemm D., Schmidt S., Höcker B. (2022) PocketOptimizer 2.0: A modular framework for computer-aided ligand-binding design Prot Sci [PubMed]
Ferruz N., Schmidt S., Höcker B. (2021) ProteinTools: a toolkit to analyze protein structures Nucleic Acids Res [PubMed]
Herud-Sikimic, Stiel A.C., Kolb M., Shanmugaratnam S., Berendzen K.W., Feldhaus C., Höcker B., Jürgens G. (2021) A biosensor for the direct visualization of auxin Nature [PubMed]
Predictive Reagent Antibody Replacement Technology (PRe-ART & PRe-ART 2T)
- together with the groups of Andreas Plückthun and Anna Hine -
In a multidisciplinary approach combining basic science, protein engineering and computational prediction, PRe-ART will generate non-antibody components which can be combined in an entirely modular fashion to create molecules that bind any peptide of choice. Such a biotechnological revolution is timely because of current failings in the reagent antibody market. More information can be found here.
Kynast J.P., Höcker B.(2023) Atligator Web: A Graphical User Interface for Analysis and Design of Protein–Peptide Interactions BioDesign Res doi: 10.34133/bdr.0011 [DOI]
Kynast J.P., Schwägerl F., Höcker B. (2022) ATLIGATOR: Editing protein interactions with an atlas-based approach Bioinformatics [PubMed]
Gisdon J.F., Kynast J.P., Ayyildiz M., Hine A.V., Plückthun A., Höcker B. (2022) Modular peptide binders - development of a predective technology as alternative for reagent antibodies. Biol Chem [PubMed]

Evolution and design of protein folds
In order to understand protein function it is important to understand how the specific and efficient contemporary enzymes evolved from less specific and less efficient precursors. The evolution of the frequently encountered (βα)8-barrel fold and its relationship to other folds such as the flavodoxin-like proteins is used to investigate this question. The enzymes HisA, HisF as well as many related (βα)8-barrel enzymes are believed to have evolved from a precursor half its size, namely from (βα)4-half-barrel domains. Database searches have revealed striking structural similarities between half-barrels and proteins adopting the flavodoxin-like (βα)5-fold. The analysis of the relationship between these structurally similar protein folds is the topic of this project. Its aim is to learn about evolutionary mechanisms that allow for the adaptation of new protein folds.
Current approaches in this project are funded by an ERC Consolidator Grant.
Webcast on this topic
based on the presentation entitled 'Engineering proteins from fold fragments' presented at the joint Biochemical Society / Protein Society Focused Meeting 'Protein engineering: new approaches and applications', held in April 2013. [BiochemSociety]
Selected References:
Ferruz N., Michel F., Lobos F., Schmidt S., Höcker B. (2021) Fuzzle 2.0: Ligand Binding in Natural Protein Building Blocks Front Mol Biosci [PubMed]
Ferruz N., Noske J., Höcker B. (2021) Protlego: A Python package for the analysis and design of chimeric proteins Bioinformatics [PubMed]
Farias-Rico, J.A., Schmidt, S. & Höcker, B. (2014) Evolutionary relationship of two ancient protein superfolds. Nature Chem Biol [PubMed]
Eisenbeis, S., Proffitt, W., Coles, M., Truffault, V., Shanmugaratnam, S., Meiler, J. & Höcker, B. (2012) The potential of fragment recombination for the rational design of proteins. J Am Chem Soc [PubMed]

Enzymatic degradation of microplastics (SFB)
- together with the group of Matthias Ullmann -
Although most plastics are very biostable, there is clear evidence that microbes can degrade these materials enzymatically. By combining different biochemical and experimental techniques with computer simulations, we want to understand which properties an enzyme must have in order to efficiently target and degrade plastics. In this respect, the recently discovered enzyme PETase, which is able to degrade PET, will serve as a model system. This enzyme is of major interest because it is structurally and functionally closely related to the enzyme group of cutinases, some of which are also able to degrade PET, albeit less efficiently. Interestingly, other cutinases are not able to do this. We are also looking for new enzymes that are able to degrade plastics such as polystyrene. More information can be found here.
Weigert S., Perez-Garcia P., Gisdon F.J., Gagsteiger A., Schweinshaut K., Ullmann, G.M., Chow J., Streit W.R., Höcker B. (2022) Investigation of the halophilic PET hydrolase PET6 from Vibrio gazogenes Prot Sci [PubMed]
Menzel T., Weigert S., Gagsteiger A., Eich Y., Sittl S., Papastavrou G., Ruckdäschel H., Altstädt V., Höcker B. (2021) Impact of Enzymatic Degradation on the Material Properties of Poly(Ethylene Terephthalate). Polymers [PubMed]
Weigert S., Gagsteiger A., Menzel T., Höcker B. (2021) A versatile assay platform for enzymatic poly(ethylene-terephthalate) degradation. Protein Eng Des Sel [PubMed]


